Problem: Your Embeddings Are a Black Box
You've got a vector database full of embeddings — but you have no idea what's actually in there. Are similar documents clustering together? Are there outliers dragging down retrieval quality? You can't answer any of this by staring at 1536-dimensional floats.
UMAP (Uniform Manifold Approximation and Projection) collapses those dimensions down to 2D or 3D while preserving local structure — so you can actually see what's going on.
You'll learn:
- How to install and configure UMAP for Python
- How to project any embedding matrix to 2D
- How to build an interactive plot with color-coded labels
- What to look for once you can see your vector space
Time: 20 min | Level: Intermediate
Why This Happens
High-dimensional vectors (256, 768, 1536 dims) are impossible to inspect directly. Most debugging workflows rely on cosine similarity queries — but those only tell you about individual pairs, not the global structure of your data.
UMAP solves this by learning a low-dimensional representation that keeps nearby points close and distant points far. Unlike PCA, it handles non-linear structure well. Unlike t-SNE, it's fast enough to run on 100k+ vectors in under a minute.
Common use cases:
- Auditing an embedding model before deploying to production
- Finding mislabeled or near-duplicate documents in your corpus
- Comparing two embedding models side-by-side
- Debugging why retrieval is returning unexpected results
Solution
Step 1: Install Dependencies
pip install umap-learn matplotlib pandas numpy
# For interactive plots (recommended)
pip install plotly
Note: The package is
umap-learn, notumap. Installingumapby mistake is one of the most common errors here.
Expected: No errors. If you hit a numba conflict, pin it:
pip install "numba>=0.56,<0.60" umap-learn
Step 2: Load Your Embeddings
UMAP expects a 2D NumPy array with shape (n_samples, n_dimensions).
import numpy as np
import pandas as pd
# Option A: Load from .npy file
embeddings = np.load("embeddings.npy") # shape: (n, dims)
# Option B: Load from a DataFrame (e.g., from a vector DB export)
df = pd.read_parquet("vectors.parquet")
embeddings = np.stack(df["embedding"].values)
# Option C: Generate on the fly with sentence-transformers
from sentence_transformers import SentenceTransformer
model = SentenceTransformer("all-MiniLM-L6-v2")
texts = ["your", "documents", "here"]
embeddings = model.encode(texts, show_progress_bar=True)
print(f"Embedding matrix: {embeddings.shape}") # e.g., (5000, 384)
Expected: A NumPy array printed as (n_samples, n_dims).
If it fails:
ValueError: setting an array element with a sequence— your embeddings aren't all the same length. Filter them first.- Memory error on large datasets — sample down to 50k rows for exploration:
embeddings = embeddings[:50000]
Step 3: Run UMAP
import umap
reducer = umap.UMAP(
n_neighbors=15, # Controls local vs global structure (5-50)
min_dist=0.1, # How tightly points cluster (0.0-1.0)
n_components=2, # Output dimensions (2 for plotting)
metric="cosine", # Use cosine for text embeddings, euclidean for images
random_state=42 # For reproducible layouts
)
projected = reducer.fit_transform(embeddings)
print(f"Projected shape: {projected.shape}") # (n_samples, 2)
Parameter guide:
n_neighbors=15— good default. Lower = tighter local clusters. Higher = more global structure visible.min_dist=0.1— lower values pack clusters tighter. Use0.0to see maximum separation.metric="cosine"— use this for text/NLP embeddings. For image embeddings,euclideanis usually better.
Expected: Runs in 10-60 seconds depending on dataset size. You'll see a progress bar from numba.
Step 4: Plot the Results
Static plot (matplotlib):
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 8))
scatter = ax.scatter(
projected[:, 0],
projected[:, 1],
c=labels, # Color by category — see below for label setup
cmap="tab20",
alpha=0.6,
s=5 # Point size — reduce for large datasets
)
ax.set_title("Vector Space (UMAP 2D Projection)")
plt.colorbar(scatter, ax=ax, label="Category")
plt.tight_layout()
plt.savefig("vector_space.png", dpi=150)
plt.show()
Interactive plot (plotly — recommended):
import plotly.express as px
plot_df = pd.DataFrame({
"x": projected[:, 0],
"y": projected[:, 1],
"label": labels, # String categories work here
"text": texts, # Hover tooltip
})
fig = px.scatter(
plot_df,
x="x", y="y",
color="label",
hover_data=["text"], # Shows the actual document on hover
title="Vector Space (UMAP 2D Projection)",
opacity=0.6,
)
fig.update_traces(marker=dict(size=4))
fig.write_html("vector_space.html") # Open in browser for full interactivity
fig.show()
Setting up labels (if you don't have them):
# From a DataFrame column
labels = df["category"].values
# From a list of strings
labels = ["finance", "tech", "health", ...]
# Numeric cluster IDs (e.g., from k-means)
from sklearn.cluster import KMeans
km = KMeans(n_clusters=10, random_state=42).fit(embeddings)
labels = km.labels_
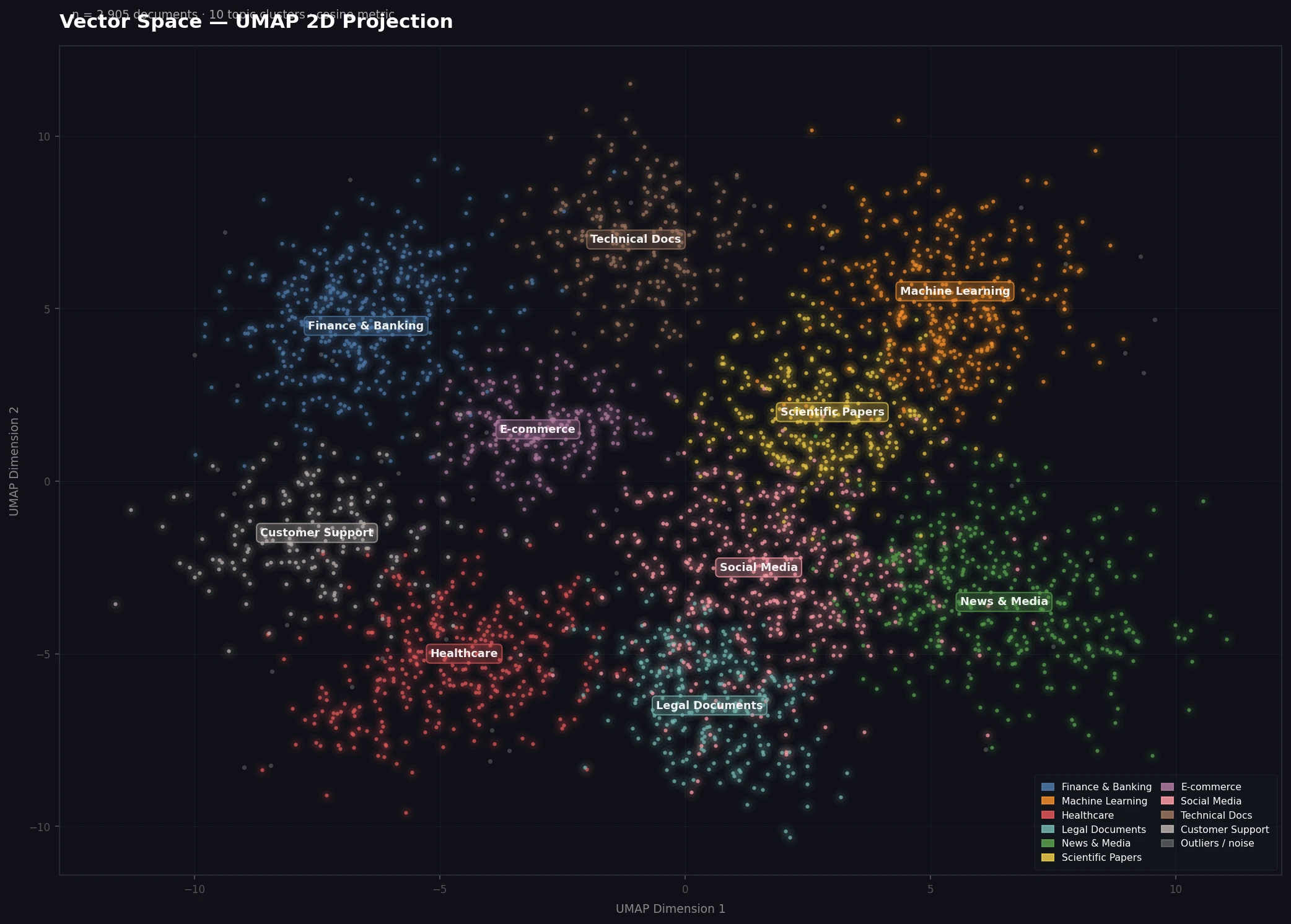 Well-separated clusters indicate your embedding model is distinguishing topics correctly
Well-separated clusters indicate your embedding model is distinguishing topics correctly
Step 5: Speed Up Large Datasets
If you have 100k+ vectors, add these settings:
reducer = umap.UMAP(
n_neighbors=15,
min_dist=0.1,
metric="cosine",
low_memory=True, # Reduces RAM at cost of speed
n_jobs=-1, # Use all CPU cores
random_state=42
)
# For very large sets: approximate nearest neighbors
# Install: pip install pynndescent
# UMAP uses this automatically when installed
For production audits on millions of vectors, project a stratified sample of 50k first to get the lay of the land.
Verification
Run this end-to-end sanity check:
# Quick smoke test with random data
import numpy as np
import umap
dummy = np.random.randn(500, 128) # 500 vectors, 128 dims
reducer = umap.UMAP(n_components=2, random_state=42)
out = reducer.fit_transform(dummy)
assert out.shape == (500, 2), "Shape mismatch"
print("UMAP working correctly.")
You should see: UMAP working correctly. in under 10 seconds.
What to Look For
Once you have the plot open, here's how to interpret what you see:
Tight, well-separated clusters — your embedding model is distinguishing categories cleanly. Retrieval should work well.
One giant blob — either your data is genuinely homogeneous, or your embedding model is too generic for this domain. Consider a domain-specific model.
Scattered outliers far from all clusters — probable mislabels, duplicates, or malformed documents. Worth inspecting those points by index:
# Find the most isolated points
from scipy.spatial.distance import cdist
center = projected.mean(axis=0)
distances = np.linalg.norm(projected - center, axis=1)
outlier_indices = np.argsort(distances)[-20:] # Top 20 outliers
print(texts[outlier_indices])
Two similar clusters close together — your categories may overlap semantically. Consider merging them or using a finer-grained model.
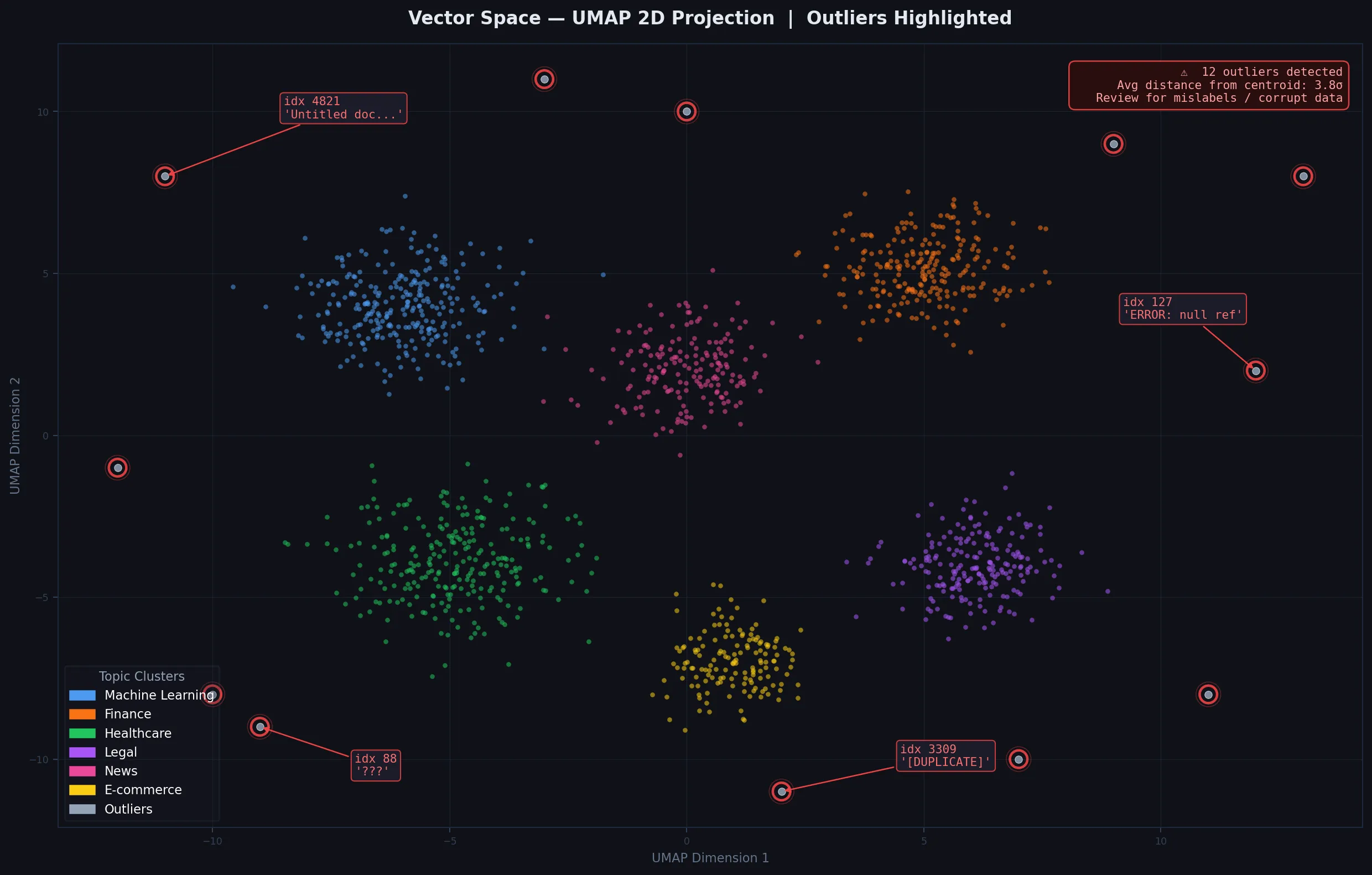 Isolated points far from clusters are worth reviewing for data quality issues
Isolated points far from clusters are worth reviewing for data quality issues
What You Learned
- UMAP projects high-dimensional embeddings to 2D while preserving local cluster structure
metric="cosine"is the right choice for text embeddings- The interactive Plotly plot is significantly more useful than static matplotlib for exploration
- Low
min_distshows tighter separation; highn_neighborsshows more global structure
Limitations to know:
- UMAP layouts are non-deterministic unless you fix
random_state— don't compare two plots without it - Distances in the 2D projection are not proportional to true cosine distances — only topology is preserved
- On GPUs, use
cuml.UMAP(RAPIDS) for 10-100x speedups on large datasets
When NOT to use this: If you need the actual low-dimensional vectors for downstream ML (not just visualization), PCA is faster and the output dimensions are interpretable.
Tested on Python 3.12, umap-learn 0.5.6, NumPy 1.26, Ubuntu 22.04 & macOS Sequoia